SpineWeb

Collaborative Platform for Research on Spine Imaging and Image Analysis
Datasets
Filed in: Main.Datasets · Modified on : Thu, 06 Dec 18
Main.Datasets History
Show minor edits - Show changes to markup
Dataset 16: 609 spinal anterior-posterior x-ray images
The dataset consists of 609 spinal anterior-posterior x-ray images. The landmarks were provided by two professional doctors in London Health Sciences Center. Each vertebra was located by four landmarks with respect to four corners. The Cobb angles were calculated using these landmarks.
Any publication resulting from any use of this database must cite the following paper:
Wu, H., Bailey, Chris., Rasoulinejad, Parham., and Li, S., 2017.Automatic landmark estimation for adolescent idiopathic scoliosis assessment using boostnet. Medical Image Computing and Computer Assisted Intervention:127-135.
To obtain, please click here
Corrections for our 2017 3D intervertebral disc localization and segmentation challenge paper
For our 2017 3D intervertebral disc localization and segmentation challenge paper, instead of computing the bidirectional Hausdorff distance, we computed the one-sided Hausdorff distance. We noticed this mistake after our paper was published. Thus, the Hausdorff distance (HD) results shown in Fig. 5 (Test1 data) and Fig. 9 (Test2 data) should be replaced with the following results.
Corrections for our 2017 3D intervertebral disc localization and segmentation challenge paper
For our 2017 3D intervertebral disc localization and segmentation challenge paper, instead of computing the bidirectional Hausdorff distance, we computed the one-sided Hausdorff distance. We noticed this mistake after our paper was published. Thus, the Hausdorff distance (HD) results shown in Fig. 5 (Test1 data) and Fig. 9 (Test2 data) should be replaced with the following results.
To obtain, please contact Jianhua Yao, Radiology and Imaging Sciences Department, Clinical Center, National Institutes of Health, USA
If you use the database for publication of any kind, please reference the following paper:
Jianhua Yao, Joseph E. Burns, Daniel Forsberg, Alexander Seitel, Abtin Rasoulian, Purang Abolmaesumi, Kerstin Hammernik, Martin Urschler, Bulat Ibragimov, Robert Korez, Tomaž Vrtovec, Isaac Castro-Mateos, Jose M. Pozo, Alejandro F. Frangi, Ronald M. Summers, Shuo Li, A Multi-center Milestone Study of Clinical Vertebral CT Segmentation, accepted for publication in Computerized Medical Imaging and Graphics, 2016.
To obtain, please click here
Dataset 15: Test set for CSI 2014 Vertebra Segmentation Challenge
This is the test data for the segmentation challenge of the CSI 2014 Workshop.
• The data sets contain 10 spine CTs acquired during daily clinical routine work in a trauma center at the Department of Radiological Sciences, University of California, Irvine, School of Medicine. Five data sets were from young adult (20-35 years old). The other five were vertebral compression fracture. • The scans cover the entire thoracic and lumbar spine. The data sets were acquired without intravenous contrast. The in-plane resolution is between 0.31 and 0.45mm and the slice thickness is 1mm or 2mm. • Image data are provided in MetaImage file format (mhd/raw). File name format is xxx.mhd.
This data is intended for research purposes, and is provided for non-commercial use only. By downloading the data, you accept the ODC Public Domain Dedication and License.
To obtain, please contact Jianhua Yao, Radiology and Imaging Sciences Department, Clinical Center, National Institutes of Health, USA
Dataset 14: IVDM3Seg – Intervertebral Disc (IVD) Localization and Segmentation from 3D Multi-Modality MR (M3) Images
An open online computational challenge in the field of spine imaging.
There are 24 3D multi-modality MRI data sets of at least 7 IVDs of the lower spine, collected from 12 subjects in two different stages in a study investigating the effect of prolonged bed rest (spaceflight simulation) on the lumbar intervertebral discs. Each subject at each stage was scanned with a 1.5-Tesla MRI scanner of Siemens using Dixon protocol. Thus, each 3D multi-modality MRI data set contains four aligned high-resolution 3D volumes: in-phase, opposed-phase, fat and water images. In total we will have 96 high resolution 3D MRI volume data. For each IVD, reference manual segmentation is provided in the form of binary mask. All images (four volumes per patient) and binary masks (one binary volume per patient) are stored in the Neuroimaging Informatics Technology Initiative (NIFTI) file format.
To obtain please contact Dr. Zheng directly. Email: guoyan.zheng@istb.unibe.ch
A detailed description of the images is found here Δ.
The Challenge is described here Δ.
The terms and conditions of usage of this dataset are here Δ
A detailed description of the images is found here.
The Challenge is described here.
The terms and conditions of usage of this dataset are here
A detailed description of the images is found here Δ.
The Challenge is described here Δ.
The terms and conditions of usage of this dataset are here Δ
A detailed description of the images is found here.
The Challenge is described here.
The terms and conditions of usage of this dataset are here
This dataset is the Testing Set A of the MICCAI 2015 Challenge “Automatic vertebral fracture analysis and identification from VFA by DXA”. It includes pairs of VFA images in two perpendicular views (lateral and anterior-posterior) for 30 subjects. They include approximately 10 subjects with compression fracture, 10 subjects with other deformations, and 10 subjects with normal shaped vertebrae. A detailed description of the images is found here. The Challenge is described here. The terms and conditions of usage of this dataset are here
This dataset is the Testing Set A of the MICCAI 2015 Challenge “Automatic vertebral fracture analysis and identification from VFA by DXA”. It includes pairs of VFA images in two perpendicular views (lateral and anterior-posterior) for 30 subjects. They include approximately 10 subjects with compression fracture, 10 subjects with other deformations, and 10 subjects with normal shaped vertebrae.
A detailed description of the images is found here.
The Challenge is described here.
The terms and conditions of usage of this dataset are here
To obtain, please click here
Dataset 12: Testing set A of the MICCAI Challenge on Vertebral Fracture Analysis
This dataset is the Testing Set A of the MICCAI 2015 Challenge “Automatic vertebral fracture analysis and identification from VFA by DXA”. It includes pairs of VFA images in two perpendicular views (lateral and anterior-posterior) for 30 subjects. They include approximately 10 subjects with compression fracture, 10 subjects with other deformations, and 10 subjects with normal shaped vertebrae. A detailed description of the images is found here. The Challenge is described here. The terms and conditions of usage of this dataset are here
Yunliang Cai, Said Osman, Manas Sharma, Mark Landis, and Shuo Li, “Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model”, IEEE Transactions on Medical Imaging, 2015.
Yunliang Cai, Said Osman, Manas Sharma, Mark Landis, and Shuo Li, “Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model”, IEEE Transactions on Medical Imaging, 2015.
Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model
More information: http://www.cg.informatik.uni-siegen.de/en/spine-segmentation-and-analysis
Further information:
Datasets came Dženan Zukić, Aleš Vlasák, Jan Egger, Daniel Hořínek, Christopher Nimsky, Andreas Kolb - Robust Detection and Segmentation for Diagnosis of Vertebral Diseases using Routine MR Images. In Computer Graphics Forum, 33(6), 2014, pages 190-204.
Dženan Zukić, Aleš Vlasák, Jan Egger, Daniel Hořínek, Christopher Nimsky, Andreas Kolb - Robust Detection and Segmentation for Diagnosis of Vertebral Diseases using Routine MR Images. In Computer Graphics Forum, 33(6), 2014, pages 190-204.
If you use these datasets in your work, it would be courteous to cite the above paper.
If you use these datasets in your work, it would be courteous to cite the above paper.
Datasets came from variety of hospitals, with a high variety (T1, T2 and TIRM sequences, and a range of TE, TR and other parameters). More details about data can be found in the following publication:
More information: http://www.cg.informatik.uni-siegen.de/en/spine-segmentation-and-analysis
http://spineweb.digitalimaginggroup.ca/images/dataset10_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset11_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset10_prev.png
Dataset 11: Spine Segmentation and Analysis
Dataset 11: High anisotropy MRIs of the lower back
To obtain, please click here
Dataset 10: Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model

To obtain, please click here
Dataset 10: Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformab le Hierarchical Model
Datasets came Dženan Zukić, Aleš Vlasák, Jan Egger, Daniel Hořínek, Christopher Nimsky, Andreas Kolb - Robust Detection and Segmentation for Diagnosis of Vertebral Diseases using Routine MR Images. In Computer Graphics Forum, 33(6), 2014, pages 190-204.
If you use these datasets in your work, it would be courteous to cite the above paper.
Dataset 11: Spine Segmentation and Analysis
Lower back pain is affecting increasing number of people with increase in office work. Authoritative diagnosis is frequently established using highly anisotropic MRI scans. This poses unique challenges. The research to tackle this was conducted between 2009 and 2014. Program code is made publicly available under an open source license, including 17 anonymized datasets with corresponding manual segmentations.
Datasets came Dženan Zukić, Aleš Vlasák, Jan Egger, Daniel Hořínek, Christopher Nimsky, Andreas Kolb - Robust Detection and Segmentation for Diagnosis of Vertebral Diseases using Routine MR Images. In Computer Graphics Forum, 33(6), 2014, pages 190-204.
If you use these datasets in your work, it would be courteous to cite the above paper.
To obtain, please click here
Datasets came Dženan Zukić, Aleš Vlasák, Jan Egger, Daniel Hořínek, Christopher Nimsky, Andreas Kolb - Robust Detection and Segmentation for Diagnosis of Vertebral Diseases using Routine MR Images. In Computer Graphics Forum, 33(6), 2014, pages 190-204.
If you use these datasets in your work, it would be courteous to cite the above paper.
http://spineweb.digitalimaginggroup.ca/images/dataset10_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset10_prev.png
To obtain, please click here
Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model
To obtain, please click this
To obtain please click [[http://spineweb.digitalimaginggroup.ca/dataset.html|here]
To obtain please click here
http://spineweb.digitalimaginggroup.ca/images/dataset10_prev.png

%width=250 ile:///Users/janemeng/Desktop/Screen%20Shot%202015-03-31%20at%209.53.42%20PM.png

%width=250 ile:///Users/janemeng/Desktop/Screen%20Shot%202015-03-31%20at%209.53.42%20PM.png

http://spineweb.digitalimaginggroup.ca/images/dataset9_prev.png
Spine structure analysis requires the understanding of vertebra locations, poses and naming in both MR and CT images. The proposed dataset aims to provide a new data reference for the increasing studies of spine recognition problems. The dataset contains MR+CT images from 20 subjects. For each vertebra, the 3D vertebra center location (at spinal cord) and the vertebra 3D orientation are manually annotated as ground truth. Any publication resulting from any use of this dataset may cite the following paper: Yunliang Cai, Said Osman, Manas Sharma, Mark Landis, and Shuo Li, “Multi-Modality Vertebra Recognition in Arbitrary Views using 3D Deformable Hierarchical Model”, IEEE Transactions on Medical Imaging, 2015
Contact: Dr. Yunliang Cai (ycai82_at_uwo.ca) or Dr. Shuo Li (slishuo_at_gmail.com)
http://spineweb.digitalimaginggroup.ca/images/dataset8_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset8_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset5_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset5_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset8_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset8_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset9_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset9_prev.png
Official website: http://lit.fe.uni-lj.si
http://spineweb.digitalimaginggroup.ca/images/dataset8_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset5_prev.png
4 segmented lumbar spine images of osteoporotic vertebral fractures (deformations).
The database consists of four CT lumbar spine images with osteoporotic vertebral fractures at one or more level (i.e. levels from L1 to L5). For each vertebra, reference manual segmentation is provided in the form of a binary mask. All images and binary masks are stored in the metaimage (MHD-RAW) format used by ITK.
The database is a preview of a larger database (of approx. 30 images) that is planned to be released by autumn 2015 as material for a tentative CSI 2016 challenge.
Any publication resulting from any use of this database must cite the following paper:
"Robert Korez, Bulat Ibragimov, Boštjan Likar, Franjo Pernuš, and Tomaž Vrtovec, “A framework for automated spine and vertebrae interpolation-based detection and model-based segmentation”, IEEE Transactions on Medical Imaging, in press, 2015 "
Contact: Robert Korez (Robert.korez@fe.uni-lj.si) or Tomaž Vrtovec (tomaz.vrtovec@fe.uni-lj.si) Laboratory of Imaging Technologies University of Ljubljana, Faculty of Electrical Engineering, Slovenia
Obtain at http://spineweb.digitalimaginggroup.ca/images/MICCAIChallengeProposal_ImagesDescription_v4.pdf
Obtain here
http://spineweb.digitalimaginggroup.ca/images/dataset9_prev.png
Obtain at http://spineweb.digitalimaginggroup.ca/images/MICCAIChallengeProposal_ImagesDescription_v4.pdf
Dataset 9: Automatic vertebral fracture analysis and identification from VFA by DXA
Osteoporotic vertebral fractures are difficult to diagnose and are often only discovered when the spine is imaged. A clinical assessment of vertebral fracture status can be performed using dual energy x-ray absorptiometry (DXA). This approach is known as vertebral fracture assessment (VFA). The datasets provided for the challenge consist of lateral and posterior-anterior (PA) scan images of the thoracolumbar spine acquired at a resolution of between 1 and 0.35 mm with Hologic Discovery A DXA scanner using the Instant Vertebral Assessment (IVA) scan option. Examples of these images (showing no vertebral fractures) are shown here (PA view (left), lateral view (right)).
In contrast with traumatic fractures, osteoporotic fractures typically affect the vertebral body. Depending on the fracture morphology, they can be classified into three types: concave, wedge and crush.
The major challenge encountered during the vertebral fracture assessment process is to determine whether deviations in the size and shape of vertebrae are due to true fractures or are simply non-fracture deformities, commonly observed in the general population.
The datasets include cases with osteoporotic vertebral fractures, cases with non-fracture deformities, and normal cases. This classification is included per vertebra and per patient.
4 segmented lumbar spine images of osteoporotic vertebral fractures (deformations).’
4 segmented lumbar spine images of osteoporotic vertebral fractures (deformations).
Attach: http://spineweb.digitalimaginggroup.ca/images/dataset7_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset7_prev.png
Attach: http://spineweb.digitalimaginggroup.ca/images/dataset7_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset7_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset7_prev.png
Dataset 8: Osteoporotic vertebral fractures
4 segmented lumbar spine images of osteoporotic vertebral fractures (deformations).’ Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
http://spineweb.digitalimaginggroup.ca/images/dataset6_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset6_prev.png
Dataset 6: Intervertebral Disc Localization and Segmentation Multi-modality MRI Spine Image Database
The database consists of multi-modality MRI spine images of 8 anonymized patients, each containing at least 7 intervertebral discs (IVDs) of the lower spine (T11 – L5). Each patient was scanned with 1.5 Tesla MRI scanner of Siemens. Dixon protocol was used to reconstruct four aligned high-resolution 3D volumes during one data acquisition (Fig. 1): in- phase, opposed-phase, fat and water images. For each IVD, reference manual segmentation is provided in the form of a binary mask. All images (four volumes per patient) and binary masks (one binary volume per patient) are stored in the Neuroimaging Informatics Technology Initiative (NIFTI) file format.
Fig. 1. The four aligned channels of a patient data (for visualization purpose, we only show the middle sagittal (mid-sagittal) slice of each channel.
http://spineweb.digitalimaginggroup.ca/images/dataset6_prev.png
Contact:
Guoyan Zheng, PD PhD Institute for Surgical Technology and Biomechanics, University of Bern, Stauffacherstrasse 78, CH-3014, Bern Switzerland
Tel: +41-31-631-5956 Email: guoyan.zheng@istb.unibe.ch
Bulat Ibragimov (bulat.ibragimov@fe.uni-lj.si) or Tomaž Vrtovec (tomaz.vrtovec@fe.uni-lj.si) Laboratory of Imaging Technologies University of Ljubljana, Faculty of Electrical Engineering, Slovenia
Bulat Ibragimov (bulat.ibragimov@fe.uni-lj.si) or Tomaž Vrtovec
(tomaz.vrtovec@fe.uni-lj.si)
Laboratory of Imaging Technologies
University of Ljubljana, Faculty of Electrical Engineering, Slovenia
Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
Dataset 5: Lumbar vertebra segmentation CT image database
The database consists of 10 CT lumbar spine images of 10 normal subjects, each containing 5 lumbar vertebrae (i.e. levels from L1 to L5). For each vertebra, reference manual segmentation is provided in the form of a binary mask. All images and binary masks are stored in the nearly raw raster data (NRRD) format.
Any publication resulting from any use of this database must cite the following paper:
Bulat Ibragimov, Boštjan Likar, Franjo Pernuš, and Tomaž Vrtovec, “Shape representation for efficient landmark-based segmentation in 3D”, IEEE Transactions on Medical Imaging, 33(4):861-874, 2014 [doi:10.1109/TMI.2013.2296976]
Contact:
Bulat Ibragimov (bulat.ibragimov@fe.uni-lj.si) or Tomaž Vrtovec (tomaz.vrtovec@fe.uni-lj.si)
Laboratory of Imaging Technologies
University of Ljubljana, Faculty of Electrical Engineering, Slovenia
Official website: http://lit.fe.uni-lj.si
Dataset 1: CVIP Spinal CT Database
Dataset 4: CVIP Spinal CT Database
Dataset 2: Cross Modality Spinal Images for Spine Workshop
Spine: 30 pairs of spinal CT and MR from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
Dataset 3: Vertebrae Localization and Identification
The database consists of spine-focused (i.e. tightly cropped) CT scans of 125 patients with varying types of pathologies. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotations of vertebrae centroids are provided. The data has been acquired at the Department of Radiology at the University of Washington.
This is the training data for the localization and identification challenge of the CSI 2014 Workshop.
Official website: http://research.microsoft.com/spine/
Contact: Ben Glocker (b.glocker@imperial.ac.uk)
Acknowledgments:
The data has been provided by the Department of Radiology at University of Washington. The public release has been supported by the Microsoft Research Connections Medical Imaging Initiative.
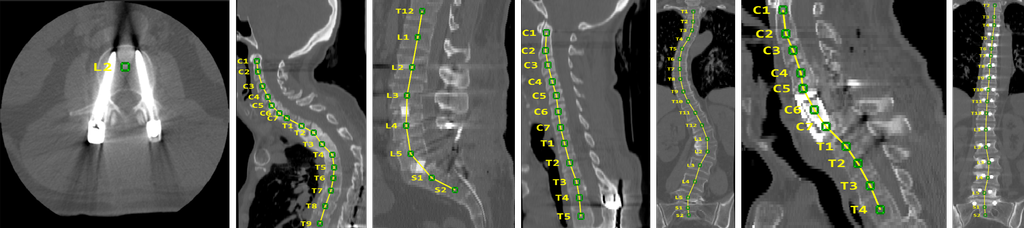
Dataset 2: Spine and Vertebrae Segmentation
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm.
This is the training data for the segmentation challenge of the CSI 2014 Workshop.
Dataset 3: Spine and Vertebrae Segmentation
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm.
This is the training data for the segmentation challenge of the CSI 2014 Workshop.
- Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
Dataset 4: Vertebrae Localization and Identification
The database consists of spine-focused (i.e. tightly cropped) CT scans of 125 patients with varying types of pathologies. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotations of vertebrae centroids are provided. The data has been acquired at the Department of Radiology at the University of Washington.
This is the training data for the localization and identification challenge of the CSI 2014 Workshop.
Official website: http://research.microsoft.com/spine/
Contact: Ben Glocker (b.glocker@imperial.ac.uk)
Acknowledgments:
The data has been provided by the Department of Radiology at University of Washington. The public release has been supported by the Microsoft Research Connections Medical Imaging Initiative.

Dataset 1: Cross Modality Spinal Images for Spine Workshop
Spine: 30 pairs of spinal CT and MR from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
- Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
- Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
Dataset 1: Cross Modality Spinal Images for Spine Workshop
Dataset 1: CVIP Spinal CT Database
The CVIP spinal CT database is provided for only research purposes.
All publications and studies that use any of the provided database must reference [1] or [2]:
[1]Melih S. Aslan, Ahmed Shalaby, and Aly A. Farag. "Clinically desired segmentation method for vertebral bodies." In Biomedical Imaging (ISBI), 2013 IEEE 10th International Symposium on, pp. 840-843. IEEE, 2013.
[2] Melih S. Aslan, Asem Ali, Ham Rara, Ben Arnold, Aly A. Farag, Rachid Fahmi, and Ping Xiang. "A novel 3d segmentation of vertebral bones from volumetric ct images using graph cuts." In Advances in Visual Computing, pp. 519-528. Springer Berlin Heidelberg, 2009.
Those who will need more would need contact with Dr. Aly A. Farag (aly.farag AT louisville DOT edu ) to fill a form for CVIP Lab.
What is given with this data:
- CT scans of 5 patients (both dicom and bmp formats)
- Ground truths: All scans have ground truths with original size (i.e., 512x512). In some data, if a VB is not fully scanned, we do not manually segment those half scans.
1 Anonbv 154_fbhs Nim=80;
2 Anonja_fbacks_25_64 Nim=64;
3 Anonsm 2355 Nim=96;
4 Ivy3_80 Nim=80;
5 MESAYUXXREM Nim=29;
http://spineweb.digitalimaginggroup.ca/images/dataset4_prev.png
Dataset 2: Cross Modality Spinal Images for Spine Workshop
Dataset 2: Spine and Vertebrae Segmentation
Dataset 3: Spine and Vertebrae Segmentation
Dataset 3: Vertebrae Localization and Identification
Dataset 4: Vertebrae Localization and Identification
Dataset 4: CVIP Spinal CT Database
The CVIP spinal CT database is provided for only research purposes.
All publications and studies that use any of the provided database must reference [1] or [2]:
[1]Melih S. Aslan, Ahmed Shalaby, and Aly A. Farag. "Clinically desired segmentation method for vertebral bodies." In Biomedical Imaging (ISBI), 2013 IEEE 10th International Symposium on, pp. 840-843. IEEE, 2013.
[2] Melih S. Aslan, Asem Ali, Ham Rara, Ben Arnold, Aly A. Farag, Rachid Fahmi, and Ping Xiang. "A novel 3d segmentation of vertebral bones from volumetric ct images using graph cuts." In Advances in Visual Computing, pp. 519-528. Springer Berlin Heidelberg, 2009.
Those who will need more would need contact with Dr. Aly A. Farag (aly.farag AT louisville DOT edu ) to fill a form for CVIP Lab.
What is given with this data:
- CT scans of 5 patients (both dicom and bmp formats)
- Ground truths: All scans have ground truths with original size (i.e., 512x512). In some data, if a VB is not fully scanned, we do not manually segment those half scans.
1 Anonbv 154_fbhs Nim=80;
2 Anonja_fbacks_25_64 Nim=64;
3 Anonsm 2355 Nim=96;
4 Ivy3_80 Nim=80;
5 MESAYUXXREM Nim=29;
http://spineweb.digitalimaginggroup.ca/images/dataset4_prev.png
%width=500pxhttp://spineweb.digitalimaginggroup.ca/images/dataset4_prev.png
http://spineweb.digitalimaginggroup.ca/images/dataset4_prev.png
%width=500pxhttp://spineweb.digitalimaginggroup.ca/images/dataset4_prev.png
1 Anonbv 154_fbhs Nim=80; 2 Anonja_fbacks_25_64 Nim=64; 3 Anonsm 2355 Nim=96; 4 Ivy3_80 Nim=80; 5 MESAYUXXREM Nim=29;
1 Anonbv 154_fbhs Nim=80;
2 Anonja_fbacks_25_64 Nim=64;
3 Anonsm 2355 Nim=96;
4 Ivy3_80 Nim=80;
5 MESAYUXXREM Nim=29;


Dataset 4: CVIP Spinal CT Database
The CVIP spinal CT database is provided for only research purposes.
All publications and studies that use any of the provided database must reference [1] or [2]:
[1]Melih S. Aslan, Ahmed Shalaby, and Aly A. Farag. "Clinically desired segmentation method for vertebral bodies." In Biomedical Imaging (ISBI), 2013 IEEE 10th International Symposium on, pp. 840-843. IEEE, 2013.
[2] Melih S. Aslan, Asem Ali, Ham Rara, Ben Arnold, Aly A. Farag, Rachid Fahmi, and Ping Xiang. "A novel 3d segmentation of vertebral bones from volumetric ct images using graph cuts." In Advances in Visual Computing, pp. 519-528. Springer Berlin Heidelberg, 2009.
Those who will need more would need contact with Dr. Aly A. Farag (aly.farag AT louisville DOT edu ) to fill a form for CVIP Lab.
What is given with this data:
- CT scans of 5 patients (both dicom and bmp formats)
- Ground truths: All scans have ground truths with original size (i.e., 512x512). In some data, if a VB is not fully scanned, we do not manually segment those half scans.
1 Anonbv 154_fbhs Nim=80; 2 Anonja_fbacks_25_64 Nim=64; 3 Anonsm 2355 Nim=96; 4 Ivy3_80 Nim=80; 5 MESAYUXXREM Nim=29;
Spine: 30 pairs of spinal CT and MRI from lumber. Each pair of CT and MRI is from same subject. Ground truth is not yet available.
Spine: 30 pairs of spinal CT and MR from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
Spine: 30 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
Spine: 30 pairs of spinal CT and MRI from lumber. Each pair of CT and MRI is from same subject. Ground truth is not yet available.
Spine: 20 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
Spine: 30 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
This is the training data for the segmentation challenge of the CSI 2014 Workshop].
This is the training data for the segmentation challenge of the CSI 2014 Workshop.
This is the training data for the localization and identification challenge of the CSI 2014 Workshop].
This is the training data for the localization and identification challenge of the CSI 2014 Workshop.
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm. More data sets will be provided for the planned MICCAI 2014 challenge on spine segmentation.
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm.
This is the training data for the segmentation challenge of the CSI 2014 Workshop].
This is the training data for the localization and identification challenge of the CSI 2014 Workshop].
(:toc anchors=invisible:)
(:toc anchors=visible:)
(:toc:)
(:toc anchors=invisible:)


Official website: http://research.microsoft.com/spine/ \\
Official website: http://research.microsoft.com/spine/ \\
Official website: http://research.microsoft.com/spine/

Acknowledgments:\\
Official website: http://research.microsoft.com/spine/ \\
The data has been provided by the Department of Radiology at University of Washington. The public release has been supported by the Microsoft Research Connections Medical Imaging Initiative.
Acknowledgments:
The data has been provided by the Department of Radiology at University of Washington. The public release has been supported by the Microsoft Research Connections Medical Imaging Initiative.

Acknowledgments
Acknowledgments:
\\
Dataset 1: Spinal Imaging and Image Analysis
Spine: 10 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available
Dataset 1: Cross Modality Spinal Images for Spine Workshop
Spine: 20 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available.
- Spine: 10 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available
Spine: 10 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available
- Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
- Cardiac: MRI from 40 subjects with LV manually segmented. Submit a request from http://digitalimaginggroup.ca/contact.php
Spinal Imaging and Image Analysis Dataset
Dataset 1: Spinal Imaging and Image Analysis
Annotated CT Database for Spine and Vertebrae Segmentation
Dataset 2: Spine and Vertebrae Segmentation
Annotated CT Database for Vertebrae Localization and Identification
Dataset 3: Vertebrae Localization and Identification
Spinal Imaging and Image Analysis Dataset
- Spine: 10 pairs of spinal CT and MRI from lumber. Each pair of CT and MR is from same subject. Ground truth is not yet available
- Cardiac: MRI from 40 subjects with LV manually segmented. Submit a request from http://digitalimaginggroup.ca/contact.php
- Obtain at http://spineweb.digitalimaginggroup.ca/dataset.html
Jianhua Yao, Joseph E. Burns, Hector Muñoz and Ronald M. Summers, Detection of Vertebral Body Fractures Based on Cortical Shell Unwrapping, International Conference on Medical Image Computing and Computer Assisted Intervention, Volume 7512, 2012, pp 509-516
Annotations
Annotations
Terms and Conditions
Terms and Conditions
Coming soon...
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm. More data sets will be provided for the planned MICCAI 2014 challenge on spine segmentation.
Data Description
- The data sets are spine CTs acquired during daily clinical routine work in a trauma center from 10 young adult (16-35 years old). The scans cover the entire thoracic and lumbar spine. The data sets were acquired without intravenous contrast. The in-plane resolution is between 0.31 and 0.45mm and the slice thickness is 1mm.
- The data have been acquired at the Department of Radiological Sciences, University of California, Irvine, School of Medicine
- Images were acquired with Philips or Siemens multidetector CT scanners.
- Image data are provided in MetaImage file format (mhd/raw). File name format is xxx.mhd.
Annotations
- In each scan, all complete thoracic and lumbar vertebrae have been semi-automatically segmented and verified.
- The manual segmentation mask is stored in MetaImage file format (mhd/raw). Each vertebra is assigned a different label. First vertebra is labeled as 100, second as 200, and so on. File name is xxx_label.mhd.
Terms and Conditions
- If you use the database for publication of any kind, please reference the following paper:
Jianhua Yao, Joseph E. Burns, Hector Muñoz and Ronald M. Summers, Detection of Vertebral Body Fractures Based on Cortical Shell Unwrapping, International Conference on Medical Image Computing and Computer Assisted Intervention, Volume 7512, 2012, pp 509-516
- This data is intended for research purposes, and is provided for non-commercial use only. By downloading the data, you accept the ODC Public Domain Dedication and License (PDDL) (http://opendatacommons.org/licenses/pddl/1.0/).
People Involved
- Jianhua Yao1 (primary contact, jyao@cc.nih.gov)
- Joseph Burns2
- Sasha Getty2
- James Stieger1
- Ronald Summers1
1. Radiology and Imaging Sciences Department, Clinical Center, National Institutes of Health, USA
2. Department of Radiological Sciences, University of California, Irvine, School of Medicine
Annotated Spine CT Database for Vertebrae Localization and Identification
Annotated CT Database for Vertebrae Localization and Identification
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm. More data sets will be provided for the planned MICCAI 2014 challenge on spine segmentation.
Coming soon
Annotated Spine CT Database for Benchmarking of Vertebrae Localization and Identification
Annotated CT Database for Spine and Vertebrae Segmentation
Segmentation is an essential step for many computational spine imaging tasks. Clinical datasets raise many difficulties for automatic methods and ground truth is often scarce. To facilitate and foster the research on this topic, we provide a database of 10 spine CT scans along with manual segmentation of the thoracic and lumbar vertebrae. The data set can be used for development, training and evaluation of spine segmentation algorithm. More data sets will be provided for the planned MICCAI 2014 challenge on spine segmentation.
Annotated Spine CT Database for Vertebrae Localization and Identification
This data release has been supported by the Microsoft Research Connections Medical Imaging Initiative.
The data has been provided by the Department of Radiology at University of Washington. The public release has been supported by the Microsoft Research Connections Medical Imaging Initiative.


Acknowledgments
This data release has been supported by the Microsoft Research Connections Medical Imaging Initiative.
Contact: Ben Glocker (b.glocker@imperial.ac.uk)
Contact: Ben Glocker (b.glocker@imperial.ac.uk) \\
Contact: Ben Glocker mailto:b.glocker@imperial.ac.uk
Contact: Ben Glocker (b.glocker@imperial.ac.uk)
Contact: Ben Glocker mailto:b.glocker@imperial.ac.uk
Official website: http://research.microsoft.com/spine/
Official website: http://research.microsoft.com/spine/

The database consists of spine-focused (i.e. tightly cropped) CT scans of adult patients (older than 18) with varying types of pathologies. The database currently consists of 125 patients. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotations of vertebrae centroids are provided.
The database consists of spine-focused (i.e. tightly cropped) CT scans of 125 patients with varying types of pathologies. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotations of vertebrae centroids are provided. The data has been acquired at the Department of Radiology at the University of Washington.
The database consists of spine-focused (i.e. tightly cropped) CT scans of adult patients (older than 18) with varying types of pathologies. The database currently consists of 125 patients. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotation of vertebrae centroids are provided.
The database consists of spine-focused (i.e. tightly cropped) CT scans of adult patients (older than 18) with varying types of pathologies. The database currently consists of 125 patients. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotations of vertebrae centroids are provided.
Annotated Spine CT Database for Benchmarking of Vertebrae Localization and Identification
The database consists of spine-focused (i.e. tightly cropped) CT scans of adult patients (older than 18) with varying types of pathologies. The database currently consists of 125 patients. For most patients, multiple scans from longitudinal examinations are available, resulting in overall 242 scans in the database. For each scan, manual annotation of vertebrae centroids are provided.
Official website: http://research.microsoft.com/spine/
Powered by PmWiki